|
|
1 year ago | |
|---|---|---|
| data/gan | 2 years ago | |
| imgs | 2 years ago | |
| networks | 3 years ago | |
| u_d | 3 years ago | |
| utils | 3 years ago | |
| .gitignore | 2 years ago | |
| LICENSE | 3 years ago | |
| README.md | 1 year ago | |
| identical_mapping.py | 3 years ago | |
| predict.py | 3 years ago | |
| requiresment.txt | 2 years ago | |
| train.py | 2 years ago | |
README.md
An Official PyTorch Implementation of “Towards Unbiased COVID-19 Lesion Localisation and Segmentation via Weakly Supervised Learning (ISBI-2021)” 
by Yang Yang, Jiancong Chen, Ruixuan Wang*, Ting Ma, Lingwei Wang, Jie Chen, Wei-Shi Zheng, Tong Zhang*.
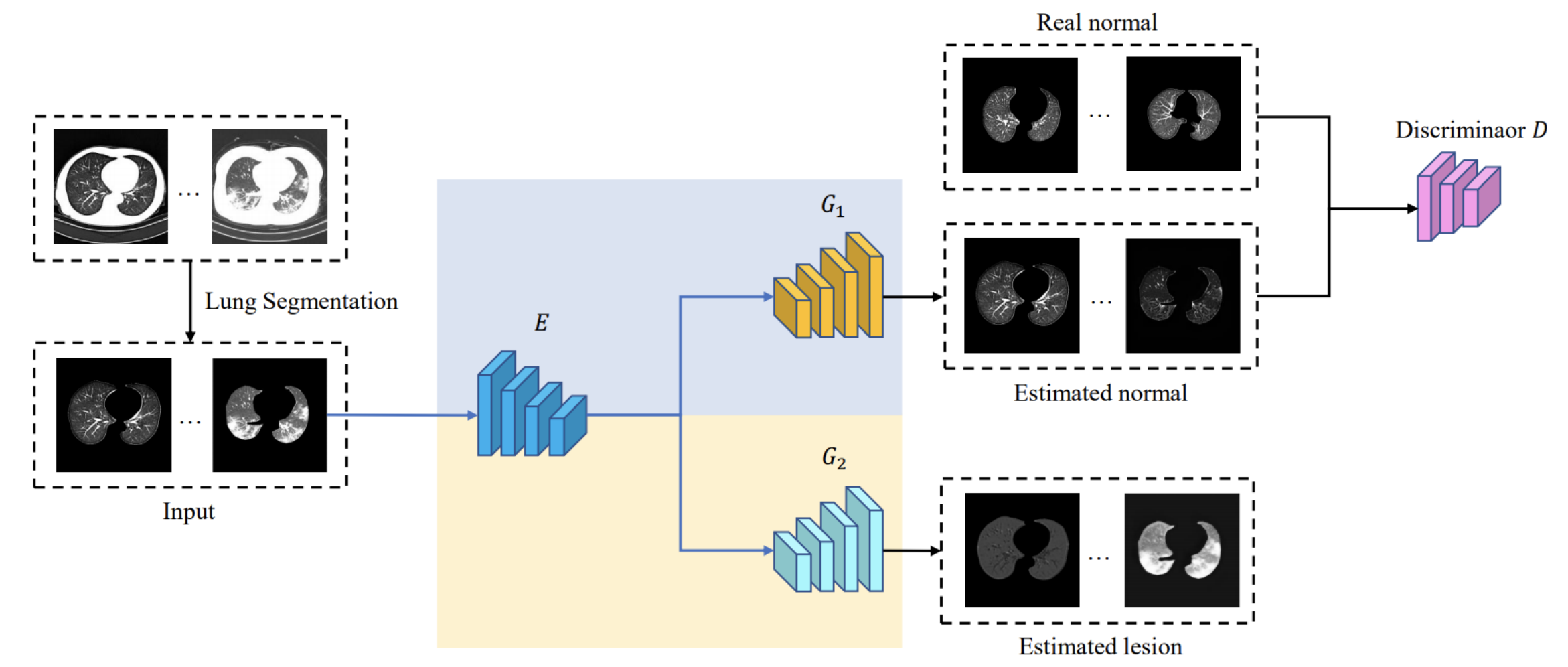
Framwork
@inproceedings{yang2021towards,
title={Towards Unbiased COVID-19 Lesion Localisation and Segmentation via Weakly Supervised Learning},
author={Yang, Yang and Chen, Jiancong and Wang, Ruixuan and Ma, Ting and Wang, Lingwei and Chen, Jie and Zheng, Wei-Shi and Zhang, Tong},
booktitle={IEEE International Symposium on Biomedical Imaging},
pages={1966--1970},
year={2021},
organization={IEEE}
}
Abstract
Despite tremendous efforts, it is very challenging to generate a robust model to assist in the accurate quantification assessment of COVID-19 on chest CT images. Due to the nature of blurred boundaries, the supervised segmentation methods usually suffer from annotation biases. To support unbiased lesion localisation and to minimise the labelling costs, we propose a data-driven framework supervised by only image level labels. The framework can explicitly separate potential lesions from original images, with the help of an generative adversarial network and a lesion-specific decoder. Experiments on two COVID-19 datasets demonstrates the effectiveness of the proposed framework and its superior performance to several existing methods.
Installation
-
Python 3.8+
-
Pytorch 1.10+
-
Torchvision 0.11.0
-
GPU, such as a 1080Ti
You can install require modules with following command.
pip install -r requirements.txt
Data
Data is stored in ./data, you can directory download it (~400M).
Training and Evaluation
You can train with following code, which save logs and models to /exp/exp-1.
python train.py -p ./exp/exp-1 -l 10 -s 0.01 -a 0.01 -g 200 -t 0 -i 15 -b 4 -e 120 --debug
Citation
If you find this project useful for your research, please cit our work use the following BibTeX entry.
@INPROCEEDINGS{9433806,
author={Yang, Yang and Chen, Jiancong and Wang, Ruixuan and Ma, Ting and Wang, Lingwei and Chen, Jie and Zheng, Wei-Shi and Zhang, Tong},
booktitle={2021 IEEE 18th International Symposium on Biomedical Imaging (ISBI)},
title={Towards Unbiased Covid-19 Lesion Localisation And Segmentation Via Weakly Supervised Learning},
year={2021},
volume={},
number={},
pages={1966-1970},
doi={10.1109/ISBI48211.2021.9433806}}
Contact
If you have any questions about this paper, welcome to email to zhangt02@pcl.ac.cn