ENGLISH | 简体中文





MindSpore SPONGE
Introduction
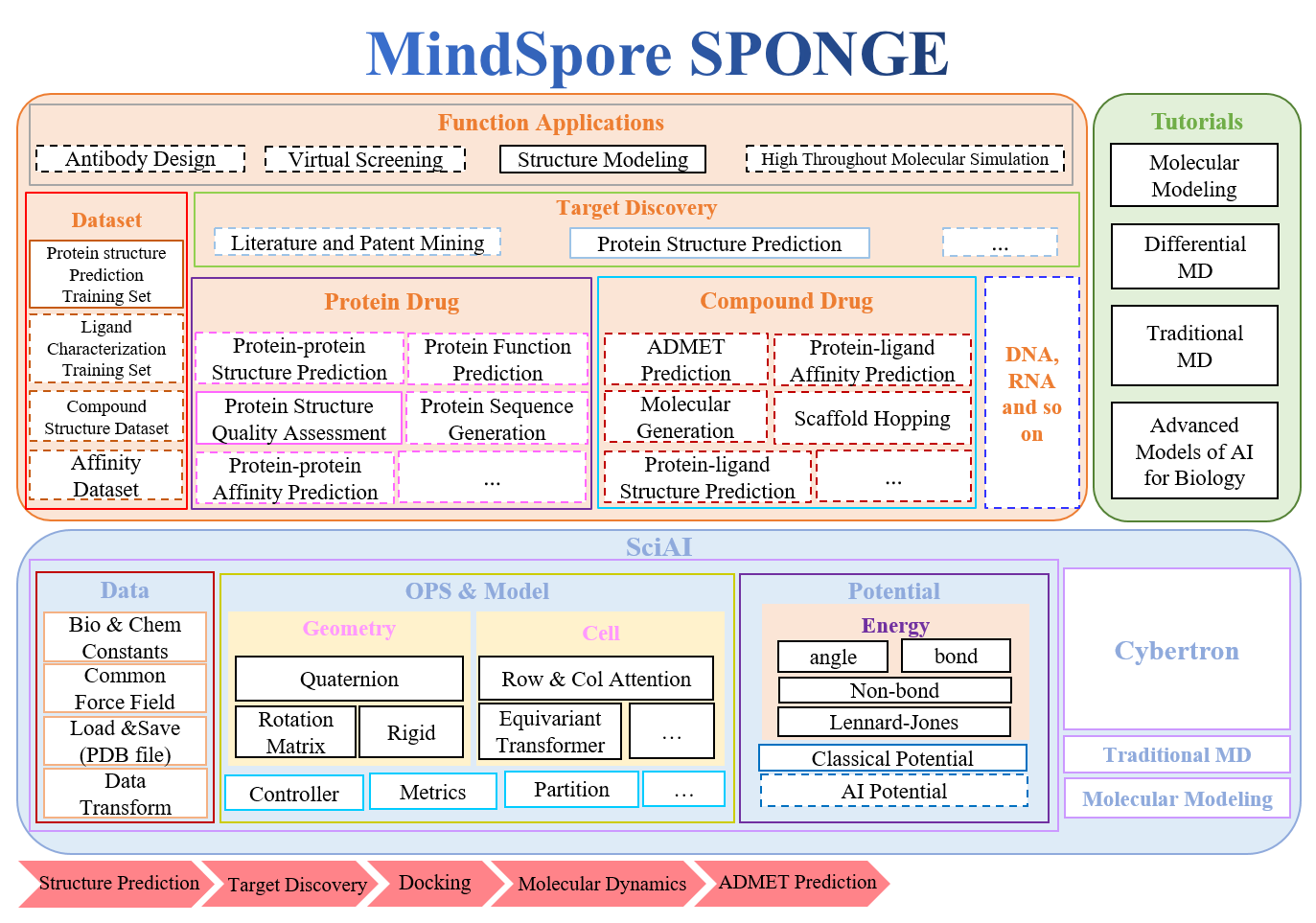
MindSpore SPONGE(Simulation Package tOwards Next GEneration molecular modelling) is a toolkit for Computational Biology based on AI framework MindSpore,which supports MD, folding and so on. It aims to provide efficient AI computational biology software for a wide range of scientific researchers, staff, teachers and students.
Latest News 📰
- 🔥
2023.12.07 The antibody design model Tiangong won the "2023 AIIA Top Ten Pioneer Application Cases of Artificial Intelligence", Related News
- 🔥
2023.11.10 MEGA-EvoGen Paper "Unsupervisedly Prompting AlphaFold2 for Accurate Few-Shot Protein Structure Prediction" is published in JCTC. Please refer to paper and code.
- 🔥
2023.8.21—2023.8.25 MindSpore SPONGE SIG Summer School is coming soon !
- 🔥 open source internship task has been released! Everyone is welcome to claim it~
- 🔥
2023.6.26 MindSPONGE Paper "Artificial Intelligence Enhanced Molecular Simulations" is published in JCTC and achieve Most Read Articles. Please refer to paper.
- 🔥
2023.5.31 Paper "Assisting and Accelerating NMR Assignment with Restrained Structure Prediction" is preprinted in arxiv, Please refer to paper and code.
2023.1.31 MindSPONGE version 1.0.0-alpha is released. The documents are available on Scientific Computing MindSPONGE module on MindSpore website2022.8.23 Paper "Few-Shot Learning of Accurate Folding Landscape for Protein Structure Prediction" is preprinted in arxiv, Please refer to paper2022.8.11—2022.8.15 MindSpore SPONGE SIG Summer School, replay2022.07.18 Paper "SPONGE: A GPU-Accelerated Molecular Dynamics Package with Enhanced Sampling and AI-Driven Algorithms"is published in Chinese Journal of Chemistry. Please refer to paper and codes2022.07.09 MEGA-Assessment wins CAMEO-QE monthly 1st2022.06.27 Paper "PSP: Million-level Protein Sequence Dataset for Protein Structure Prediction" is preprinted in arxiv. Please refer to paper and codes.2022.04.21 MEGA-Fold wins CAMEO-3D monthly 1st. Related News
Quick Start
Protein Multimer Structure Prediction
import os
import stat
import pickle
from mindsponge import PipeLine
from mindsponge.common.protein import to_pdb_v2, from_prediction_v2
cmd = "wget https://download.mindspore.cn/mindscience/mindsponge/Multimer/examples/6T36.pkl"
os.system(cmd)
pipe = PipeLine(name="Multimer")
pipe.set_device_id(0)
pipe.initialize("predict_256")
pipe.model.from_pretrained()
f = open("./6T36.pkl", "rb")
raw_feature = pickle.load(f)
f.close()
final_atom_positions, final_atom_mask, confidence, b_factors = pipe.predict(raw_feature)
unrelaxed_protein = from_prediction_v2(final_atom_positions,
final_atom_mask,
raw_feature["aatype"],
raw_feature["residue_index"],
b_factors,
raw_feature["asym_id"],
False)
pdb_file = to_pdb_v2(unrelaxed_protein)
os.makedirs('./result/', exist_ok=True)
os_flags = os.O_RDWR | os.O_CREAT
os_modes = stat.S_IRWXU
pdb_path = './result/unrelaxed_6T36.pdb'
with os.fdopen(os.open(pdb_path, os_flags, os_modes), 'w') as fout:
fout.write(pdb_file)
print("confidence:", confidence)

More Cases:👀
Installation
Hardware
| Hardware |
OS |
Status |
| Ascend 910 |
Ubuntu-x86 |
✔️ |
|
Ubuntu-aarch64 |
✔️ |
|
EulerOS-aarch64 |
✔️ |
|
CentOS-x86 |
✔️ |
|
CentOS-aarch64 |
✔️ |
| GPU CUDA 10.1 |
Ubuntu-x86 |
✔️ |
pip install
pip install https://ms-release.obs.cn-north-4.myhuaweicloud.com/2.2.1/MindScience/mindsponge/ascend/aarch64/mindsponge_ascend-1.0.0rc2-py3-none-any.whl
pip install https://ms-release.obs.cn-north-4.myhuaweicloud.com/2.2.1/MindScience/mindsponge/gpu/x86_64/cuda-10.1/mindsponge_gpu-1.0.0rc2-py3-none-any.whl
The version of mindsponge installed by pip corresponds to the r0.5 branch code. The code can be downloaded using the following instruct.
git clone -b r0.5 https://gitee.com/mindspore/mindscience.git
source code install
git clone https://gitee.com/mindspore/mindscience.git
cd {PATH}/mindscience/MindSPONGE
bash build.sh -e ascend
export CUDA_PATH={your_cuda_path}
bash build.sh -e gpu -j32
cd {PATH}/mindscience/MindSPONGE/output
pip install mindsponge_ascend*.whl # Ascend
pip install mindsponge-gpu*.whl # GPU
API
For details about MindSPONGE APIs, please refer to API pages.
CO-CHAIR
Special Interesting Group 🏠
MindSpore SPONGE SIG (Special Interesting Group) is a team composed of a group of people who are interested and have a mission to make achievements in the field of AI × biological computing.
MindSpore SPONGE SIG group provides efficient and easy-to-use AI computational biology software for researchers, teachers and students, and provides a platform for people with strong abilities or strong interests in this field to communicate and cooperate together.
At present, the SIG group has six core teachers. After members joining the SIG group, our teachers will lead the team to carry out scientific research and develop the software function development. Of course, members are also welcome to do research on their own topics using MindSPONGE.
In the SIG group, we will hold various activities, including summer school, public lecture, technology communication meeting and other large-scale activities. Small-scale activities like weekly meetings, blogs writing will also be held in the group. By joining the activities, there will be lots of chances to communicate with our experts. During the summer school program ended on August 15th, we invited 13 teachers to have a five-day lecture mainly including three themes of MindSpore basics, molecular dynamics and advanced AI × Science courses. You can get the replay here.
In the SIG group, we will also release the public intelligence task and open source internship task, welcome everyone to claim it.
If you want to join us and become a member of our group, please send your resume to liushuo65@huawei.com, we are always looking forward to your arrival.
Core Contributor 🧑🤝🧑
Cooperative Partner
Contribution Guide
License
Apache License 2.0