Are you sure you want to delete this task? Once this task is deleted, it cannot be recovered.
|
|
1 year ago | |
|---|---|---|
| models | 1 year ago | |
| utils | 1 year ago | |
| LICENSE | 1 year ago | |
| README-zh.md | 1 year ago | |
| README.md | 1 year ago | |
| data_function.py | 1 year ago | |
| data_preprocessing.py | 1 year ago | |
| hparam.py | 1 year ago | |
| loss_function.py | 1 year ago | |
| main.py | 1 year ago | |
| metrics.py | 1 year ago | |
| show_data.py | 1 year ago | |
| train.py | 1 year ago | |
README.md
Pytorch Medical Segmentation
Read Chinese Introduction:Here!
Notes
We are planning a major update to the code in the near future, so if you have any suggestions, please feel free to email me or mention them in the issue.
Recent Updates
- 2021.1.8 The train and test codes are released.
- 2021.2.6 A bug in dice was fixed with the help of Shanshan Li.
- 2021.2.24 A video tutorial was released(https://www.bilibili.com/video/BV1gp4y1H7kq/).
- 2021.5.16 A bug in Unet3D implement was fixed.
- 2021.5.16 The metric code is released.
- 2021.6.24 All parameters can be adjusted in hparam.py.
- 2021.7.7 Now you can refer medical classification in Pytorch-Medical-Classification
- 2022.5.15 Now you can refer semi-supervised learning on medical segmentation in SSL-For-Medical-Segmentation
- 2022.5.17 We update the training and inference code and fix some bugs.
Requirements
- pytorch1.7
- torchio<=0.18.20
- python>=3.6
Notice
- You can modify hparam.py to determine whether 2D or 3D segmentation and whether multicategorization is possible.
- We provide algorithms for almost all 2D and 3D segmentation.
- This repository is compatible with almost all medical data formats(e.g. nii.gz, nii, mhd, nrrd, ...), by modifying fold_arch in hparam.py of the config. I would like you to convert both the source and label images to the same type before using them, where labels are marked with 1, not 255.
- If you want to use a multi-category program, please modify the corresponding codes by yourself. I cannot identify your specific categories.
- Whether in 2D or 3D, this project is processed using patch. Therefore, images do not have to be strictly the same size. In 2D, however, you should set the patch large enough.
Prepare Your Dataset
Example1
if your source dataset is :
source_dataset
├── source_1.mhd
├── source_1.zraw
├── source_2.mhd
├── source_2.zraw
├── source_3.mhd
├── source_3.zraw
├── source_4.mhd
├── source_4.zraw
└── ...
and your label dataset is :
label_dataset
├── label_1.mhd
├── label_1.zraw
├── label_2.mhd
├── label_2.zraw
├── label_3.mhd
├── label_3.zraw
├── label_4.mhd
├── label_4.zraw
└── ...
then your should modify fold_arch as *.mhd, source_train_dir as source_dataset and label_train_dir as label_dataset in hparam.py
Example2
if your source dataset is :
source_dataset
├── 1
├── source_1.mhd
├── source_1.zraw
├── 2
├── source_2.mhd
├── source_2.zraw
├── 3
├── source_3.mhd
├── source_3.zraw
├── 4
├── source_4.mhd
├── source_4.zraw
└── ...
and your label dataset is :
label_dataset
├── 1
├── label_1.mhd
├── label_1.zraw
├── 2
├── label_2.mhd
├── label_2.zraw
├── 3
├── label_3.mhd
├── label_3.zraw
├── 4
├── label_4.mhd
├── label_4.zraw
└── ...
then your should modify fold_arch as */*.mhd, source_train_dir as source_dataset and label_train_dir as label_dataset in hparam.py
Training
- without pretrained-model
set hparam.train_or_test to 'train'
python main.py
- with pretrained-model
set hparam.train_or_test to 'train'
python main.py -k True
Inference
- testing
set hparam.train_or_test to 'test'
python main.py
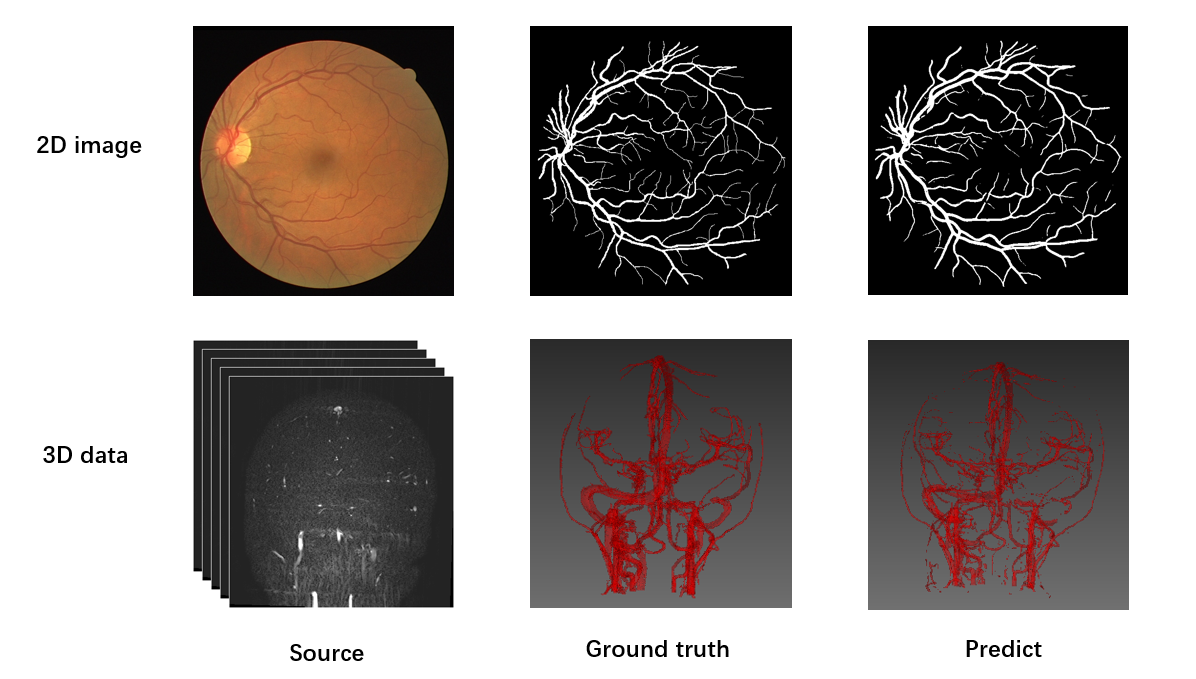
Examples
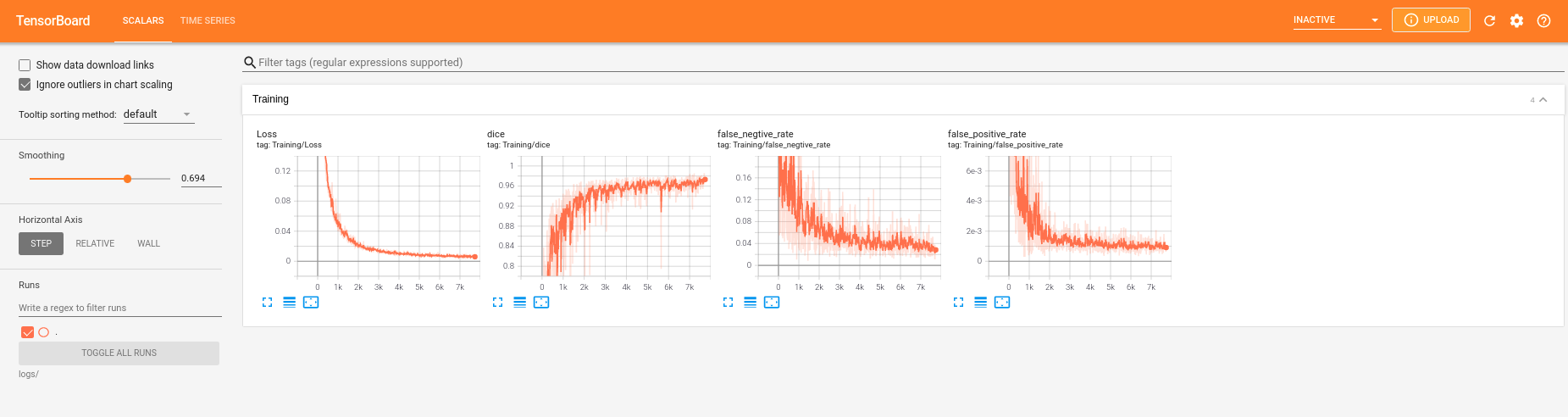
Tutorials
Done
Network
- 2D
- 3D
Metric
TODO
By The Way
This project is not perfect and there are still many problems. If you are using this project and would like to give the author some feedbacks, you can send Me an email.
Acknowledgements
This repository is an unoffical PyTorch implementation of Medical segmentation in 3D and 2D and highly based on MedicalZooPytorch and torchio. Thank you for the above repo. The project is done with the supervisions of Prof. Ruoxiu Xiao and Dr. Cheng Chen. Thank you to Youming Zhang, Daiheng Gao, Jie Zhang, Xing Tao, Weili Jiang and Shanshan Li for all the help I received.
Related works
If this code is helpful for you, you can cite these for us. Thank you.
[1] Chen C, Zhou K. An Effective Deep Neural Network for Lung Lesions Segmentation from COVID-19 CT Images[J]. IEEE Transactions on Industrial Informatics, 2021.
[2] Chen C, Zhang T, et al. Pathological lung segmentation in chest CT images based on improved random walker[J]. Computer Methods and Programs in Biomedicine, 2021, 200: 105864.